- Research
- Open access
- Published:
Automated reasoning using abduction for interpretation of medical signals
Journal of Biomedical Semantics volume 5, Article number: 35 (2014)
Abstract
This paper proposes an approach to leverage upon existing ontologies in order to automate the annotation of time series medical data. The annotation is achieved by an abductive reasoner using parsimonious covering theorem in order to determine the best explanation or annotation for specific user defined events in the data. The novelty of this approach resides in part by the system’s flexibility in how events are defined by users and later detected by the system. This is achieved via the use of different ontologies which find relations between medical, lexical and numerical concepts. A second contribution resides in the application of an abductive reasoner which uses the online and existing ontologies to provide annotations. The proposed method is evaluated on datasets collected from ICU patients and the generated annotations are compared against those given by medical experts.
Introduction
Medical monitoring of patients is becoming increasingly device supported and thus large volumes of high frequency data are generated from sensors that monitor physiological parameters. While the use of such technologies enables a continuous monitoring, the complexity and amount of data creates a challenge for the medical staff to provide interpretations. Furthermore, such interpretations may be complex as sensor data is inherently uncertain, there may exist interdependencies between physical parameters, and the data is voluminous and multivariate [1, 2].
Automated analysis and mining techniques have the potential to support the medical staff in the interpretation of the data. For time series data analysis this implies a need for proper annotation of the signals with domain dependent knowledge in order to facilitate decision making and eventual diagnosis. The output generated by the algorithms should ideally provide information that is compatible with the knowledge and the terms used by health practitioners. In data-driven approaches [3] the labelling of data is limited to those pre-defined by the engineers implementing the algorithms. On the other hand, knowledge driven approaches offer the possibility to more explicitly model the relations between higher level concepts and data. However, these techniques e.g. rule based methods, also require significant manual effort to encode domain knowledge.
At the same time, the amount of structured knowledge in the medical domain is rapidly increasing due in part by the Linked Data model. This model which is based on the RDF model [4] allows bodies of knowledge that are independently structured to be directly interlinked without any further customization efforts. For example, NCBO BioPortal[5] as a repository of biomedical ontologies contains more than 300 ontologies holding about 5 millions classes that cover medical concepts including the causes and symptoms of diseases. The rise of large and shared machine processable knowledge repositories provides an opportunity to automate the utilisation of information.
In this paper, we propose a system which is able to receive as input time series signals and generate as output an annotation of these signals. Domain knowledge is inputted into the system in a flexible manner allowing the practitioners to express freely the terms and thresholds that are relevant for a particular physiological parameter i.e. an event. To enable flexibility, these expressions are connected to a number of ontologies containing relations between concepts expressed by the practitioner and observations measured by the various sensors. The ontologies used are the Symptom ontology as one of the ontologies in BioPortal [5, 6], WordNet[7] and the Semantic Sensor Network (SSN) ontologya. The symptom ontology provides the medical terms and definitions defined as concepts in a hierarchy of subsumption relations which are used in the annotations of the sensor data. The WordNet ontology which consists of a lexical database of the English language enables finding relations between the concepts in BioPortal and those defined by the practitioner. The SSN ontology is used to link the specific sensors to physiological parameters, and provide a standardized representation of sensors, observations and related concepts.
The reasoning process used in this paper which finds the relations contained in the different ontologies, is abductive. Abductive reasoning is chosen as it is non-monotonic and thus differs from deductive reasoning in that a logically certain conclusion is not guaranteed. Rather, abductive reasoning infers the best possible explanation given a set of observations. Techniques such as Parsimonious Covering Theory (PCT) or diagnostic reasoners which are abductive are often used in the medical domain [8] as they promote explicitation, and can contend with uncertainty by assessing the likelihood that a specific hypothesis entails a given conclusion [9].
This paper whose main focus is more on the reasoning method and its scalability and less on the auxiliary techniques such as Natural Language Processing (NLP) used, evaluates the use of existing ontologies and abductive reasoning to annotate sensor data from ICU patients. One benchmark dataset provided for use in 1994 AI in Medicine symposium submissions [10] and one dataset collected at a local hospital (Section ‘DataSets’) are used in the experimental analysis. The annotations generated by the proposed approach are compared against the annotations made by experts. Also, the complexity of the reasoning method is evaluated.
The paper begins with a description of related works in Section ‘Related work’. The Linked Data model and Abductive Reasoning are then shortly introduced in Section ‘Background’. We explain the details of the framework in Section ‘Method’ and then discuss the results of the reasoner and evaluate the framework’s output in Section ‘Results and discussion’. The paper ends with the conclusion and discussion in Section ‘Conclusion’.
Related work
In the literature, research whose goal is to use knowledge driven methods to annotate time series data is found in various fields in artificial intelligence that include sensor data enrichment [11, 12], data stream annotation [13], symbol grounding [14, 15], and semantic perception [16]. Such works share the common feature where symbolic knowledge is integrated to the numeric data processing. Often high level symbolic knowledge is manually encoded based on the requirements of the problem rather than (re)using existing knowledge already modelled in e.g., ontologies (RDF graph model). For example, [16] and [17] have proposed reasoning techniques based on abductive reasoning for data stream annotation using manually encoded knowledge. These works including [18] implemented in OWL use PCT for inferring the best possible explanation. However, the reasoner is restricted to generate explanations with only one cause. The work presented in [19] implements an automated reasoning which is similar to our work in the sense that the knowledge base consists of a RDF/OWL ontology. However, in our work, we propose an automated reasoning over external ontologies modelled by different experts. Furthermore, the PCT based reasoner in our work overcomes the constraint of providing an explanation containing more than one cause for the observations. This approach builds upon previous work [20] and has formalized the reasoning process and extended the experimental evaluation.
Background
In this section we introduce preliminary features of the Linked Data model and abductive reasoning.
Linked data
Exploiting human knowledge for commonsense and automated reasoning has always been a challenge. The fast-growing Webb which has traditionally been populated with HTML documents is known as the biggest repository of human knowledge in different domains. However, despite the fact that contents of this repository are accessible in the form of pages, due to the lack of semantic interconnection among them, it is impossible for an artificial agent to retrieve a specific concept. Therefore, the first step towards automatically using the content of Web pages is structuring these contents so that they become interlinked and can be queried in different levels of abstraction.
Linked data which refers to a set of structured data, namely global data space, has become a paradigm providing the transition from document oriented Web into a web of interlinked data [21]. According to this paradigm, unstructured information represented in web pages is mapped into the RDF graph which is understood as a set of subject-predicate-object triples, [4]. Given  as a set of dereferenceable URIsc and
as a set of dereferenceable URIsc and  as a set of literals such as numbers or strings, the aforementioned RDF triple is defined as . In other words, all subjects and predicates are URIs and objects are either a URI or a literal value. Similarly, stating the set , where
as a set of literals such as numbers or strings, the aforementioned RDF triple is defined as . In other words, all subjects and predicates are URIs and objects are either a URI or a literal value. Similarly, stating the set , where  as a set of variables is ranging over , we can redefine the triple
as a set of variables is ranging over , we can redefine the triple  as an element of the query set
as an element of the query set  . More specifically, instead of feeding search engines with search terms, it is possible to fetch the desired set of triples by writing a query which is equivalent to the finite set of triples Q. Eventually, an answer for this query is simply achieved by binding variables of the query triples into .
. More specifically, instead of feeding search engines with search terms, it is possible to fetch the desired set of triples by writing a query which is equivalent to the finite set of triples Q. Eventually, an answer for this query is simply achieved by binding variables of the query triples into .
Different languages such as RDFS and OWL complying with the Linked Data model, provide different levels of expressivity. Regardless of the implementation language, however, it is the uniformity and the integrability features of the Linked Data model that make the integration of different linked datasets straightforward.
However, despite its unified structure, there are number of issues with linked data that pose a challenge for automated reasoning [22]. For instance, in order to query large size linked datad, the query process needs to deal with the problem of localizing relevant parts in linked data.
In this paper, a biomedical repository called BioPortal [5] is used. Using a similar data model as the Linked Data model, BioPortal contains more than 300 ontologies ranging in subjects from anatomy, phenotype description, to health [6]. Further details about dealing with the aforementioned issue of size are discussed in Section ‘Hypothesis extraction’.
Abductive reasoning with PCT
Reasoning processes are categorized into two main groups, monotonic and non-monotonic reasoning. Monotonic reasoning including deductive reasoning implies that inferring a new piece of knowledge does not change the set of already known information. Non-monotonic reasoning, on the other hand, states that adding more knowledge can invalidate current conclusions. In diagnostic medical procedure where symptoms of a disease gradually emerge, monotonic reasoning due to the permanence of its results, are less favourable. Since all the symptoms of a disease do not occur at a same time, the reasoner needs to be able to deal with incomplete data throughout the reasoning process. Incompleteness may also extend to the high level models e.g. ontologies which may also be dynamically changing. A non-monotonic reasoning process whose set of answers can later be updated is therefore useful in domains such as medicine and industrial diagnosis process [23].
There are different models of non-monotonic reasoning such as default reasoning, autoepistemic logic, belief revision and abductive reasoning [23]. In this work, we selected abductive reasoning with the ability of deriving the best (most likely) explanations out of known facts. Abduction as the backbone of commonsense reasoning and has increasingly been applied in diagnosis systems (medical domains) [24]. Diagnostic reasoning in particular, is based on abductive logic and represents the knowledge within a network of causal associations. The “hypothesis-and-test” approach of diagnosis reasoning shows its non-monotonic behaviour where the set of plausible causes of the observed behaviour can change whenever the observation set extends. Parsimonious Covering Theory (PCT) is a model of diagnostic reasoning [8] used in this work.
PCT formalization is based on set theory and is defined within a quadruple , where  is the set of all observations which are either qualitative or quantitative objects;
is the set of all observations which are either qualitative or quantitative objects;  states the set of all manifestations (events) detected over observations; subsequently,
states the set of all manifestations (events) detected over observations; subsequently,  contains all hypotheses defined as possible causes that are in relations with expected events. Finally,
contains all hypotheses defined as possible causes that are in relations with expected events. Finally,  is the solution set indicating inferred explanations for items of
is the solution set indicating inferred explanations for items of  . More specifically, the inference process is about drawing as an explanation for elements of . However, the formalization is not complete in that it does not formalize the “best” explanation. For this, PCT suggests various criteria to select the final result set
. More specifically, the inference process is about drawing as an explanation for elements of . However, the formalization is not complete in that it does not formalize the “best” explanation. For this, PCT suggests various criteria to select the final result set  . Two widely used criteria are:
. Two widely used criteria are:
-
Set covering criterion is defined as a property of a function f which is assumed to be a mapping from a subset of
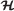 (set of all hypotheses) to a subset of
(set of all hypotheses) to a subset of  (set of manifestations) so that
(set of manifestations) so that  is a possible cause for . An accepted conclusion w.r.t the set-covering criterion is set such that .
is a possible cause for . An accepted conclusion w.r.t the set-covering criterion is set such that . -
Minimum cardinality criterion is concerned about the cardinality of the solution set. According to this criterion,
 as a subset of
as a subset of  is chosen as the solution set if for all other “covering” subsets of
is chosen as the solution set if for all other “covering” subsets of 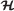 , namely
, namely 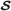 , .
, .
As previously mentioned, PCT is based on set theory. The eventual explanation is a subset of the of the Hypothesis set for which aforementioned criteria hold. Selecting a subset poses an issue of the time complexity. Consequently, there are a number of techniques that address computational factors for making abductive reasoning NP-Hard [25]. For instance, applying constraints that reduce the composite hypothesis set size as well as ruling out criteria-violating candidates (and their super classes) from the power set, can reduce the time complexity. Techniques used in this work are further discussed in Section ‘Reasoner’.
Method
The reasoner depicted in Figure 1 receives two primary inputs, Hypothesis ( ), and Manifestation (
), and Manifestation ( ) which are separately provided by the HypothesisExtractor and the ManifestationExtractor processes, respectively. The output of the reasoner is called Explanation (
) which are separately provided by the HypothesisExtractor and the ManifestationExtractor processes, respectively. The output of the reasoner is called Explanation ( ). Each component feeding the reasoner contains several modules that collaborate with ontologies including the SSN ontology and the WordNet ontology.
). Each component feeding the reasoner contains several modules that collaborate with ontologies including the SSN ontology and the WordNet ontology.
Considering the PCT quadruple explained in Section ‘Abductive reasoning with PCT’, we then follow the reasoning process of the framework by mapping the main elements of PCT into outputs of different components.
Configuration
The framework illustrated in Figure 1 is based on a configuration file which is filled by the expert of the domain. The configuration file contains details of (possible) behaviours of signals in which the expert is interested to monitor. To illustrate the method in the paper, we will use a running example of configuration files shown in Figure 2, 3 and 4. For instance, Figure 2 is about a situation where the expert is interested to observe the “heart rate”, “amount of oxygen saturation” and “blood pressure”. There is also a section in the configuration file in which the expert, by setting a range of values, can specify a significant behaviour for physiological terms.
Configuration file sample II (related to the same infant in Figure 2 ).
The SSN ontology is populated only with the contents of the configuration file. There is an equivalent class or property in SSN, for each item (key/value pair) mentioned in the configuration file. The value of a key in the file is used as a name of a class in SSN. Given FeatureOfInterest and Property as concepts defined in SSN and the function valueOf(key) which returns the value of a key in the configuration file, the SSN ontology is populated as follows:
For example, the SSN ontology populated with the content of Figure 2 will contain the following axioms:
The configuration file allows expert to enter values which denote either a normal or an abnormal behaviour in signals. For example, in Figure 2 and 4 we can find the definition of abnormal and normal behaviours, respectively. In the experimental validation in Section ‘Results’, we show that the eventual explanations are not literally dependent on the content of the file. More specifically, the signal explanation process results in same interpretation for variations of terms used by the expert.
Hypothesis extraction
According to PCT, the Hypothesis set is defined as a set of facts that represent relations between expected events and their causes. The SemanticAnalyser module (Figure 1) initializes the process resulting in the Hypothesis set. This module collaborating with public ontologies is responsible for retrieving a hierarchy of related concepts formatted in RDF/OWL.
Before going to the details of the Hypothesis Extraction, we first explain how we deal with localizing the relevant parts in Bioportal. The goal of the system is annotating medical signals that contain abnormal behaviours, (i.e., symptoms of diseases). SemanticAnalyser queries for the term “symptom” in the NCBO BioPortal. The results of this query is 21 records out of which 15 items belong to the Symptom ontology, as a sub ontology in BioPortal. Therefore, due to its high rank, the Symptom ontology is chosen as a reference ontology.
The symptom ontology illustrated in Figure 5, has been modelled to capture signs and symptoms of diseases and provides well-categorized medical symptoms in terms of body part names. Due to its structure, the symptom ontology is only used for retrieving the hierarchy of symptom concepts modelled based on subsumption relations. Running the following SPARQL querye, the SemanticAnalyser module retrieves a hierarchy of symptoms in terms of subclasses of the “symptom” class:

SemanticAnalyser, then searches through the set of symptom classes in order to select relevant symptoms. The relevant symptoms are those ones that are related to parts of the body (“feature_of_interest”) observed by sensors e.g., “heart” and “blood” in case of the configuration file in Figure 2, or “heart”, “blood” and “respiratory” system in case of the configuration file in Figure 4. In order to find the relevant symptoms, each symptom type passes the two phases of tokenizingf and stemmingg. As shown in Figure 1, the SemanticAnalyser module uses the WordNet ontology that contains synonym/pertainymh set of words and acquires the synonym/pertainym set of each token of a symptom type. Consequently, each symptom type (split into its tokens) is assigned with multiple synonym/pertainym lists corresponding to its tokens. The number of times that each physiological parameter (the “feature_of_interest” value) appears in the synonym set of each token is counted. Finally, a symptom type whose tokens have the highest total count is chosen as the top candidate which has the highest similarity to the “feature_of_interest”. Table 1 shows all categorized symptom types along with the body parts’ name for different configuration files. For example, the “cardiovascular system symptoms” is chosen due to its highest relevance to the “heart” as a “feature_of_ interest”.
The final Causes set shown in Figure 1 is the union of all subclasses of the candidate symptom types returned per each “feature_of_interest”. In Table 2, cause items as the output of the SemanticAnalyser module are listed. The first 62 items and the total 89 items are considered as causes related to configurations in Figure 2 and Figure 4, respectively. As we see in Table 2, each cause can be a single term (e.g., hypoxemia) or a combination of terms (e.g., atrial fibrillation). The definition of each single cause term is retrieved from either the Symptom ontology or the WordNet ontology (in case the former returns nothing) to be replaced with the cause item.

The Hypothesis set is generated by the SignalMapper module. The SignalMapper takes as input the set of causes. It selects a subset of these causes based on parameters mentioned in the configuration file. Specifically it looks at the terms used to define behaviours. For example, possible behaviours for a specific signal are defined as “fast”, “slow” and “irregular”. The SignalMapper concatenates the values of behaviour, feature of interest and property to generate a list of phrases such as “irregular heart rate”, “low oxygen saturation” (see Table 3). For those configurations where the expert states the normal behaviour, e.g., Figure 4, the term “not” is added in front of the concatenated phrase, e.g. “not normal respiratory rate”. For phrases preceded by “not” an antonym set is retrieved from WordNet.
As the next step towards generating the Hypothesis set, the SignalMapper process builds an n×m similarity matrix S, where n and m are the number of cause items and the number of possible behaviours, respectively. The similarity matrix S which is initialized to zero, will hold the similarity values between elements of these two lists (Algorithm 1). For calculating the similarity values, the cause items need to be grammatically analysedi. For instance, for each cause item, grammatical roles of its terms such as noun (“NN”) or adjective (“JJ”) are identified. All causes will therefore have their own grammatical tree by running the grammatical analysing process over rows of the matrix. In order to set the value of element si,j of matrix S, the process first needs to generate the grammatical structure tree of the ith cause and then to check whether this cause is related to an behaviour j. For this, all adjectives (“JJ”) with their own substantives (“NN”) of the ith cause item are retrieved. Each substantive (called noun1) is also checked to see if it is related (e.g., via a preposition or a connective) to another noun (called noun2). If such a combination is found in a cause item, at the next step, the synonym/pertainym sets of the adjective, noun1 and noun2 are also retrieved to be checked against the column side items. The value of si,j increases if the following three conditions are met: (SynSet(K) refers to the synonym/pertainym set of term K, c j refers to the jth column and r i refers to the ith row)
Figures 6 and 7 illustrate two samples of a grammatical structure tree for two causes and their relations with two behaviours. The matrix element referring to “arrhythmia” and “irregular heart rate” will be set to 1 due to the matching terms found between them (Figure 6). Likewise, after running the process of Algorithm 1, the matrix element referring to “tachypnea” and “not normal respiratory rate” is set to 1 (Figure 7).
After calculating the elements’ value of the matrix S, the SignalMapper chooses non zero elements showing a relation between causes and behaviours. The Hypothesis set ( ) as the first input of the reasoner (Figure 1) is created by pairs of row-column items of non-zero elements in the matrix S. Table 4(a) and Table 4(b) partially show two retrieved Hypothesis sets based on the two configurations in Figure 2 and Figure 4, respectively.
) as the first input of the reasoner (Figure 1) is created by pairs of row-column items of non-zero elements in the matrix S. Table 4(a) and Table 4(b) partially show two retrieved Hypothesis sets based on the two configurations in Figure 2 and Figure 4, respectively.
Manifestation extraction
The ManifestationExtractor component is responsible for the signal analysis process. This component contains a module called SignalAnalyser (Figure 1) which performs the event detection process. Using the SSN ontology which is only populated with the configuration information, the Signal Analyser detects those parts of signals that contain an abnormal behaviour mentioned in the configuration. An event (or an abnormal behaviour detected in a signal) is defined based on threshold values set by the expert of the domain according to sampling rate of signals and the patient profile (age, gender, etc.). For example, in Figure 2, the “Behaviour” section related to “heart” shows the range of “heart rate” values as “ <157 A N D >175” which is set by the expert to monitor the situation of an infant to detect an “irregular” heart behaviour. The expert would enter different values in case of an adult patient. For instance, the upper bound of the “slow heart rate” for an infant is set to 157 (Figure 2) while the same behaviour for an adult patient is set to 60 (Figure 4).
The applied data analysis method divides signals into several segments. A segment is created based on the number of events (set as threshold values defining a numeric range) detected in each signal. The division process is done within an iterative process which looks for events in each signal and determines a set of temporal intervals in which a number of events are included. The iterative process starts by creating a temporal segment in the first signal whose length is set based on the minimum required number of events in the signal. More precisely, the starting time point of the initial segment is the same as that of the signal, and its ending point is when the minimum required number of events in this signal has been met. Detecting a new event affects the size and the number of created intervals. The iteration ends whenever the size of intervals do not change. At the end, these intervals are considered as segments. The reasoner will then be applied on each segment separately. Therefore, the threshold values set by the expert enables him/her to have some segments in which, for example, one signal has no event while the others do.
Although the data analysis method can affect the eventual interpretation results, it is the representation technique which, in this work, is at focus. In Section ‘Results’, examples of the threshold values set for a configuration is given.
The output of the ManifestationExtractor component is a set of Manifestations ( ) at each segment, which is a list of time points at which events are detected. The Manifestation set is the second input of the reasoner (Figure 1).
) at each segment, which is a list of time points at which events are detected. The Manifestation set is the second input of the reasoner (Figure 1).
Reasoner
The reasoner module is based on Parsimonious Covering Theory (PCT) as an abductive reasoning method whose basis is on the set theory. The main feature of this reasoner is finding the best possible Explanations ( ) for the set of Manifestations (
) for the set of Manifestations ( ) detected at each segment of signals. More precisely, given the Hypothesis set (
) detected at each segment of signals. More precisely, given the Hypothesis set ( ) which is the set of the cause/abnormal_behaviour pairs, the reasoner calculates the power setk of the causes set. Final Explanations are those members of the power set (or subsets of the causes set) which do not violate the reasoner’s principles.
) which is the set of the cause/abnormal_behaviour pairs, the reasoner calculates the power setk of the causes set. Final Explanations are those members of the power set (or subsets of the causes set) which do not violate the reasoner’s principles.
The principles of the reasoner are defined within two criteria: Covering and Minimality. According to the first criterion shown in (1), the reasoner nominates those subsets of the causes set ( ) that are related to all Manifestations. In other words, the covering set indicates a set of subsets of causes with the aforementioned specification. Moreover, the concern of the minimality criterion (2) is the size of the selected subset. Complying with aforementioned criteria, the reasoner finds the best possible explanations which are those covering subsets of the causes (as part of the Hypothesis set) that are minimal in terms of the cardinality. Algorithm 2 shows the details of the reasoner.
) that are related to all Manifestations. In other words, the covering set indicates a set of subsets of causes with the aforementioned specification. Moreover, the concern of the minimality criterion (2) is the size of the selected subset. Complying with aforementioned criteria, the reasoner finds the best possible explanations which are those covering subsets of the causes (as part of the Hypothesis set) that are minimal in terms of the cardinality. Algorithm 2 shows the details of the reasoner.

The reasoning complexity, due to the power set calculation, grows exponentially w.r.t the number of causes. In order to reduce the size of the power set, two steps indicated in Algorithm 2 are applied. The first step filters the set of causes by removing those causes that are not listed in pairs of the Hypothesis set. At the second step, super classes are removed for elements of the power set where the minimality criterion is violated.
The output of the reasoner is the set of Explanations for observations.
Results and discussion
DataSets
In order to evaluate the framework, we use two different sets of multivariate medical data. The first dataset contains 12-hours of time-series data from a set of medical sensors measuring heart rate, arterial pressure, and arterial oxygen saturation of an infant in an Intensive Caring Unit (ICU). This patient is suffering from several diseases, namely “multiple liver abscesses”, “portal hypertension” and “E. Coli sepsis”, used as the ground truth for the evaluation of the final explanations suggested by the reasoner. This package of data is the ICU data package provided for use in 1994 AI in Medicine symposium submissions [10]. The second dataset also contains multivariate data from three sensors measuring heart rate, respiratory rate and blood pressure of an adult patient in a Critical Caring Unit (CCU) who is suffering from “congestive heart failure (CHF)”. This package is provided by the caring unit section of a hospitall.
Results
In this section, we discuss about the experiments which are based on two different configurations and two different datasets. The first experiment is related to the configurations in Figure 2 and the infant patient data introduced above. The second experiment is based on the configurations set in Figure 4 and the adult patient data. Finally, the scalability of the reasoner is also evaluated base on different configuration parameters such as: number of “feature_of_interests” ( ), size of the Causes set (), number of abnormal behaviours (
), size of the Causes set (), number of abnormal behaviours ( ) and distinct number of causes in the Hypothesis set ().
) and distinct number of causes in the Hypothesis set ().
Experiment I
Figure 2 shows the configurations used in this experiment for monitoring the “heart” and “blood” situation of a patient. The properties of interest are “rate” (rate of heart), “pressure” (pressure of blood) and “oxygen” (amount of oxygen in blood). As mentioned in Section ‘Hypothesis extraction’, in order to find the relevant symptoms, each symptom type listed in Table 1, is assigned with the synonym/pertainym list of its tokens. For example, the set of tokens of the first symptom types (“abdominal symptoms”) is [abdomen, symptom ]whose elements are assigned to their synonym/pertainym list: abdomen ↦ { venter, stomach, belly}symptom ↦ { indication, evidence, gesture, mark, point,...} Since there is no match between items of the above lists and the two physiological parameters (heart and blood), the value of the “abdominal symptoms” item is set to zero. However, the 8th item, “cardiovascular system symptoms”, is tokenized as [cardiovascular, system, symptom]. Focusing on the first token, the synonym/pertainym list is: cardiovascular ↦ {cardiac, heart}The score of the item “cardiovascular system symptoms”, related to the “heart”, hence, increases to 1. The “hemic system symptoms” item, in a same way, gets 1 score since the pertainym of “hemic” is the term “blood”. Therefore, the selected symptoms indicated in Table 1 are those ones that are related to the “cardiovascular system” and “hemic system” symptoms due to their highest similarity values to the “feature_of_interests” set in the configuration file.
The list of 62 cause items () which are subclasses of the selected symptom types (“cardiovascular system symptoms” and “hemic system symptoms”) are only partially shown in Table 2. Furthermore, the list of all possible behaviours mentioned in the configuration file (Figure 2), that created by the SignalMapper module, is depicted in Table 3(a). As we can see, the process of concatenating “behaviour”, “feature_of_interest” and “property” values results in 7 phrases indicating different behaviours (). For example, the first item, “slow heart rate” is generated by concatenating the term “slow” as “behaviour”, the term “heart” as “feature_of_interest” and the term rate as “property”.
In order to achieve the Hypothesis set ( ) whose elements are the pairs of cause / abnormal_behaviour, the SignalMapper process, at its next step, creates a 62×7 similarity matrix S initialized to zero. The updated value of the element si,j will indicate the relation between the ith cause and the jth behaviour. As mentioned before, for each cause item whose definitions has been retrieved from the symptom or the WordNet ontology, a grammatical structure tree holding the grammatical role of each term in the sentence, is generated. Finding the similarity between causes and abnormal behaviours implies a need for checking if a similar phrase to an abnormal behaviour is detected within a cause item (Algorithm 1). For example, the first cause item (Table 2), arrhythmia, is defined as “an abnormal rate of muscle contractions in the heart”. As we see in the grammatical tree of this cause illustrated in Figure 6, there is an adjective (“abnormal”) whose substantive (“rate”) is also related to a noun, “heart” (via a preposition, “in”). This cause item is found similar to the third behaviour (“irregular heart rate”) since:
) whose elements are the pairs of cause / abnormal_behaviour, the SignalMapper process, at its next step, creates a 62×7 similarity matrix S initialized to zero. The updated value of the element si,j will indicate the relation between the ith cause and the jth behaviour. As mentioned before, for each cause item whose definitions has been retrieved from the symptom or the WordNet ontology, a grammatical structure tree holding the grammatical role of each term in the sentence, is generated. Finding the similarity between causes and abnormal behaviours implies a need for checking if a similar phrase to an abnormal behaviour is detected within a cause item (Algorithm 1). For example, the first cause item (Table 2), arrhythmia, is defined as “an abnormal rate of muscle contractions in the heart”. As we see in the grammatical tree of this cause illustrated in Figure 6, there is an adjective (“abnormal”) whose substantive (“rate”) is also related to a noun, “heart” (via a preposition, “in”). This cause item is found similar to the third behaviour (“irregular heart rate”) since:
Therefore, the element s1,3 is set to 1. Following Algorithm 1, the similarity matrix S will finally contain 18 non-zero values referring to 18 pairs cause/abnormal_behaviour that creates the Hypothesis set () (Table 4(a)). Counting the number of causes, we find 11 distinct items out of 18 in this list (). Therefore, during the reasoning process, where the power set of the causes set is generated, the reasoner needs to deal with the power set with the size of 211.
The SignalAnalyser detects abnormal behaviours of data and represents them as items of the Manifestation set ( ) for each segment of data. The applied data analysis method divides signals into several segments which as explained in Section ‘Manifestation extraction’, are defined based on the desired number of events at each signal as well as the sampling rate of the signal. Figure 8 shows three signals related to the configurations in Figure 2. The threshold value for the Arterial Pressure Signal has been set as “ 25≤n≤60” meaning that a segment needs to have at least 25 and at most 60 arterial pressure events. Similarly, the threshold values for the Arterial O2 Saturation and the Heart rate are “ 2≤n≤15” and “ 5≤n≤20”, respectively. According to these threshold values, signals in Figure 8 are divided into 3 segments.
) for each segment of data. The applied data analysis method divides signals into several segments which as explained in Section ‘Manifestation extraction’, are defined based on the desired number of events at each signal as well as the sampling rate of the signal. Figure 8 shows three signals related to the configurations in Figure 2. The threshold value for the Arterial Pressure Signal has been set as “ 25≤n≤60” meaning that a segment needs to have at least 25 and at most 60 arterial pressure events. Similarly, the threshold values for the Arterial O2 Saturation and the Heart rate are “ 2≤n≤15” and “ 5≤n≤20”, respectively. According to these threshold values, signals in Figure 8 are divided into 3 segments.
Given the two sets Hypothesis ( ) and Manifestation (
) and Manifestation ( ), the reasoner separately provides inferred Explanations for each segment shown in Table 5. For the patient of the first dataset, 6 distinct diseases (explanations) have been found (Table 6(a)). By calculating the probability of occurrence for each disease, the soundness of the reasoner outputs is evaluated. The Occurrence probability is defined as the ratio of the number of times a disease has been seen to the number of different explanations observed for a segmentm. According to Table 6(a), the first (hypertension) and the forth (Septic Shock) items are matched with the diseases mentioned in the patient profile (“portal hypertension” and “E. Coli sepsis”) with the probability of 100% and 33%, respectively. In addition, other items which are discovered by the reasoner but are not mentioned in the patient profile such as “tachycardia” and “hypertension” are in the literature considered as a sign of “Sepsis” [26]. Therefore, if we also count these combinations as sepsis, as shown in Table 6(b), the true positive diseases are the two first ones in the ordered list. The false negative case which exists in the patient profile but has not been inferred by the reasoner is “liver abscesses”. This liver dependent disease to be diagnosed, most likely requires other types of sensors information in order to be detected.
), the reasoner separately provides inferred Explanations for each segment shown in Table 5. For the patient of the first dataset, 6 distinct diseases (explanations) have been found (Table 6(a)). By calculating the probability of occurrence for each disease, the soundness of the reasoner outputs is evaluated. The Occurrence probability is defined as the ratio of the number of times a disease has been seen to the number of different explanations observed for a segmentm. According to Table 6(a), the first (hypertension) and the forth (Septic Shock) items are matched with the diseases mentioned in the patient profile (“portal hypertension” and “E. Coli sepsis”) with the probability of 100% and 33%, respectively. In addition, other items which are discovered by the reasoner but are not mentioned in the patient profile such as “tachycardia” and “hypertension” are in the literature considered as a sign of “Sepsis” [26]. Therefore, if we also count these combinations as sepsis, as shown in Table 6(b), the true positive diseases are the two first ones in the ordered list. The false negative case which exists in the patient profile but has not been inferred by the reasoner is “liver abscesses”. This liver dependent disease to be diagnosed, most likely requires other types of sensors information in order to be detected.
Experiment II
In this section, we continue the experiments with the second dataset and present results of the reasoner for situations where the expert uses the negation concept in the configuration file. As mentioned in Figure 4, the expert decided to monitor the heart rate, the blood pressure and the respiratory rate of the patient. Before going to the details, we examine the results of the HypothesisExtraction component for this case.
Candidate symptoms for the second dataset in Table 1 are “cardiovascular system”, “hemic system” and “respiratory system” symptoms. The entire subclasses of these three concepts in the Synonym ontology contain 89 causes () shown in Table 2. Moreover, for configurations in Figure 4, there are 6 possible abnormal behaviours () (see Table 3(b)). One of these items, “not normal respiratory rate”, is the phrase with negation for which the antonym set rather than the synonym set is retrieved from the WordNet ontology. SignalMapper, then, creates a 89×6 similarity matrix in order to prepare the Hypothesis set. Table 4(b) shows 21 relations () out of which 17 cause items () are distinct. Therefore, the reasoner has only to deal with 217 elements of the power set.
Due to the threshold values set for the segmentation process, the signals which are the results of 4 days of observation with the sampling rate of once per hour, is divided into 1 segment. Shown in Figure 9, the threshold values for the heart rate, respiratory rate and blood pressure are set as, “ 1≤n≤25”, “ 50≤n≤70” and “ 30≤n≤55”, respectively. The inferred Explanations are shown in Table 7.
It is worth mentioning that for the first dataset, since the cardinality of all inferred Explanations at each segment were the same (3 items for each explanation), we did not consider the minimality criterion. However, for the second dataset, since the reasoner results in explanations with different sizes, the evaluation will be different. As shown in Table 7, the first two explanations holds the minimality criteria of the reasoner, “heart failure” and “dyspnea”. The first one is matched with CHF, the disease the patient is suffering from. Furthermore, the second one, dyspnea, is considered as a main sign of heart failure in the literature [27].
Experiment III
The purpose of the following experiment is to examine the performance of the reasoner given various inputs. For example, given a larger hypothesis set, the reasoner spends more time on the processing of the power set calculation. In the following, we momentarily disregard the time for segmentation of the signals (as this is independent of the configurations) and we represent the reasoning time for different configurations in order to study the scalability of the reasoner and the impact of the parameters in a configuration file on the reasoning performance.
Recall that the final explanation is retrieved from the Hypothesis set ( ) which is as such extracted from the Cause set (
) which is as such extracted from the Cause set ( ). As said in Section ‘Hypothesis extraction’, the cause items are the union of the subclasses of the candidate symptom types. The candidate symptom types are also chosen based on the “feature_of_interest” parameters mentioned in a configuration. For instance, in the first experiment (Section ‘Experiment I’), due to the 2 mentioned “feature_of_interests” in the configuration file (), there were finally 2 symptom types chosen. Since, the number of subclasses for each symptom type is not really specified, we consider it as a constant value for all types of symptoms. Therefore, the number of symptom types which is equivalent to the number of “feature_of_interests” (
). As said in Section ‘Hypothesis extraction’, the cause items are the union of the subclasses of the candidate symptom types. The candidate symptom types are also chosen based on the “feature_of_interest” parameters mentioned in a configuration. For instance, in the first experiment (Section ‘Experiment I’), due to the 2 mentioned “feature_of_interests” in the configuration file (), there were finally 2 symptom types chosen. Since, the number of subclasses for each symptom type is not really specified, we consider it as a constant value for all types of symptoms. Therefore, the number of symptom types which is equivalent to the number of “feature_of_interests” ( ) indicated in the configuration, is considered as a significant parameter which affects the cardinality of the Cause set (). The greater the parameter
) indicated in the configuration, is considered as a significant parameter which affects the cardinality of the Cause set (). The greater the parameter  , the larger the value of . In experiment I: , ,In experimentII: , ,
, the larger the value of . In experiment I: , ,In experimentII: , ,
Since the input of the reasoning process is the Hypothesis set which is extracted from the Cause set, we focus on parameters affecting the distinct number of causes in the Hypothesis set (). The first parameter, is the size of the Cause set () which is also dependent on the  parameter. Another parameter influencing , is the number of behaviours (
parameter. Another parameter influencing , is the number of behaviours ( ).
).
In Table 8, we listed the measured reasoning time (in milliseconds)n for different configurations. The information of each row in Table 8 belongs to a configuration which is accumulated with a new configuration for its next row. In the following the summary of four configurations which are accumulated in order are given:

The first row and first configuration uses only one feature_of_interest () and the number of causes retrieved from the symptom ontology is (Table 2). The distinct number of causes in the Hypotheses is and is based on the 3 possible behaviours (). The reasoning time for calculating the power set of causes in the Hypothesis set is 1 ms. However, since the generation and the filtering process of the similarity matrix is necessary to reach to the final set, we consider the last column of the table as the final reasoning time (19 ms) which is the summation of both the similarity matrix calculation time and the reasoning time and by increasing the parameter  in the second row of Table 2 () the growth of the number of behaviours (), we see the total reasoning time also increased to 54 ms. In order to see the effect of the parameter
in the second row of Table 2 () the growth of the number of behaviours (), we see the total reasoning time also increased to 54 ms. In order to see the effect of the parameter  , we keep the same “feature_of_interests” in the third row ( and therefore ). By adding the third configuration, the only parameter changes is the number of behaviours (), which results in a much longer reasoning time (2172 ms). Although the parameter
, we keep the same “feature_of_interests” in the third row ( and therefore ). By adding the third configuration, the only parameter changes is the number of behaviours (), which results in a much longer reasoning time (2172 ms). Although the parameter  influences the reasoning time, the effect of the parameter
influences the reasoning time, the effect of the parameter  on the reasoning process is stronger.
on the reasoning process is stronger.
The reasoning process due to the techniques explained in Algorithm 2 (such as filtering the cause items based on their relations with events), is much more efficient than a pure calculation of the power set of the Cause set. Nevertheless, it still needs to deal with the power set calculation for a smaller size of causes in the Hypothesis set, explaining an exponential trend in computation time. Therefore, the system configurations for higher scales matters. For instance, behaviours allow the system to reduce the number of causes which are not relevant and results in a smaller size of  . At the same time, however, the higher number of behaviours enables the system to accept more cause items during the similarity matrix filtering process, which results in a bigger size of
. At the same time, however, the higher number of behaviours enables the system to accept more cause items during the similarity matrix filtering process, which results in a bigger size of  and consequently a higher reasoning time. The number of behaviours given in the configuration file is therefore the most influential parameter in the reasoning time. In summary, according to the computational time represented in Table 8, the user in order to have a reasonable computational time, is recommended not to define more than 3 behaviours for each property of a feature_of_interest in a configuration file.
and consequently a higher reasoning time. The number of behaviours given in the configuration file is therefore the most influential parameter in the reasoning time. In summary, according to the computational time represented in Table 8, the user in order to have a reasonable computational time, is recommended not to define more than 3 behaviours for each property of a feature_of_interest in a configuration file.
Although the intensive care units (ICUs) depending on the patient situation or medical specialty are divided into several parts such as medical intensive care unit (MICU), surgical intensive care unit (SICU), etc., there are common equipments in terms of monitoring critical physiological parameters [28]. For instance, instant monitoring of pulse oximetry, arterial blood pressure, oxygenation saturation, temperature along with using ventilators assisting the respiratory systems are done by common wired sensors used in any care units of emergency cases. Considering the typical monitoring sensors in hospitals’ care units, the computational time of our approach applied on other real world scenarios with in average 4 sensors and 3 general behaviours would be the same as what we discussed above.
Conclusion
In this paper, we have presented a framework which is able to annotate medical sensor data with labels containing probable causes pertaining to sensor events. This framework reduces the probability of losing the relevant causes by retrieving a wide possibility of causes which are related to sensor data. At the same time, by pruning the retrieved concepts (removing irrelevant causes w.r.t the probable events), the complexity of the reasoning process is reduced.
The primary motivation to the presented work is having the data annotation process that is as automated as possible. The process uses manually created configuration file which is filled by the expert of the domain and is based on events which are likely to occur. Although the process of generating explanations of the data is dependent upon the content of the configuration file, the expert is free to populate this file using his/her own words. In other words, the eventual explanations, due to synonyms of terms considered throughout the interpretation process, are literally (but not conceptually) independent of terms used by the expert. Certain limitations in the system include the level of complexity of the user defined configurations. In addition, we chose to populate the SSN ontology with classes so as to provide the opportunity of a better classifications of relevant classes for future purposes. For example, by creating the two classes Heart and Cardiac system as the subclasses of the feature_of_interest class, the system will be able to, for some purposes in future, create a “owl:sameas” properties between them to introduce them as equivalent classes.
Furthermore, as discussed in Section ‘Experiment III’, the user of the system needs to consider the limitation in number of abnormal behaviour defined in the configuration to avoid the time complexity of the reasoner to increase. In addition, the filtering process in similarity matrix, where the relevant causes are chosen based on their grammatical structure, can be further extended towards considering complicated situations that may be found in English definition of a cause.
Although the use of the symptom ontology is limited to the retrieval of subclasses, still, the existence of this ontology with its well-categorized structure was a positive feature of the medical repository which provided readable categories of symptoms in terms of different parts of the body. In order to extend the framework to be applicable to other domains (e.g., Meteorology or Geography), such a general ontology related to the domain is necessary. For this reason, the medical domain is the more promising application domain for this approach. Considering the requirements of this framework in terms of the structure of knowledge, along with the reasoning issues over linked data such as data inconsistency or redundancy may help to efficiently develop and populate linked data for different domains.
Endnotes
a This ontology developed by the W3C Semantic Sensor Networks Incubator Group (SSN-XG) describes sensors, observations, and related concepts [29].
b The size of the Web is 3.32 billion sites [30].
c URIs return contents of a resource that they identify.
dhttp://datahub.io/group/lodcloud(over 31 billion triples),
e The last access date of the Bioportal’s SPARQL endpoint (http://sparql.bioontology.org) is on 27 th July 2014
f The process of splitting a sequence of strings into its elements (tokens or words).
g The process of reducing inflected words to their stem, base or root form.
h In WordNet 2.1 OWL, pertain is a property between two WordSense concepts that indicates the relevant term for a word [7]
i In this work we used StanfordParser [31] to analyse phrases or sentences.
j It is useful to recall that the jth column refers to abnormal (or “not” + normal) behaviour that is composed of a “behaviour” (as an adjective), a “feature_of_interest” (as a noun) and a “property” (as a noun).
k The power set of a set is the set of all its subsets.
l Due to the ethical concerns about the patient’s privacy we received this dataset as an anonymous patient profile.
m Since all three segments are the same, the occurrence probability can be calculated for one segment and its values can be generalized.
n The computational time has been done on a computer which has an Intel(R) Core(TM) i7-2620M CPU (2.70GHz), 64 bit, 4 cores,4 MB for the cache memory),12 GB memory, and Linux kernel 3.8.0-44-generic.
References
Salatian A, Adepoju F:Towards an icu clinical decision support system using data wavelets. IJCA Special Issue on Systems and Data Processing (ICISD). Published by Foundation of Computer Science. 2011, 37-43.
Sukuvaara TI, Sydanmaa ME, Nieminen HO, Heikela A, Koski EMJ:Object-oriented implementation of an architecture for patient monitoring. IEEE Eng Med Biol. 1993, 12 (4): 69-81.
Jensen PB, Jensen LJ, Brunak S:Mining electronic health records: towards better research applications and clinical care. Nat Rev Genet. 2012, 13 (6): 395-405. 10.1038/nrg3208.
Heath T, Bizer C: Linked Data: Evolving the Web Into a Global Data Space. 2011, San Rafael: Morgan & Claypool
Musen MA, Noy NF, Shah NH, Whetzel PL, Chute CG, Storey M-AD, Smith B:The national center for biomedical ontology. JAMIA. 2012, 19 (2): 190-195.
Salvadores M, Alexander PR, Musen MA, Noy NF:Bioportal as a dataset of linked biomedical ontologies and terminologies in rdf. Semantic Web. 2013, 4 (3): 277-284.
Huang X-X, Zhou C-L:An owl-based wordnet lexical ontology. J Zhejiang University - Sci A. 2007, 8 (6): 864-870. 10.1631/jzus.2007.A0864.
Reggia JA, Peng Y:Modeling diagnostic reasoning: a summary of parsimonious covering theory. Comput Methods Prog Biomed. 1987, 25: 125-134. 10.1016/0169-2607(87)90048-4.
JØsang A:Abductive reasoning with uncertainty. Proceedings of the International Conference on Information Processing and Management of Uncertainty (IPMU). 2008,
Bache K, Lichman M: UCI Machine Learning Repository. [http://archive.ics.uci.edu/ml],. 2013, Irvine, CA: University of California, School of Information and Computer Science
Moraru A, Mladenic D:A framework for semantic enrichment of sensor data. J Comput Information Technol(CIT). 2012, 20 (3): 167-173.
De Mel G, Pham T, Damarla T, Vasconcelos W, Norman T:Semantically enriched data for effective sensor data fusion. SPIE Defense, Security, and Sensing. International Society for Optics and Photonics Volume 8047. 2011,
Wei W, Barnaghi P:Semantic annotation and reasoning for sensor data. Proceedings of the 4th European Conference on Smart Sensing and Context. 2009, Springer Berlin Heidelberg, 66-76.
Coradeschi S, Loutfi A, Wrede B:A short review of symbol grounding in robotic and intelligent systems. KI – Künstliche Intelligenz. 2013, 27 (2): 129-136. 10.1007/s13218-013-0247-2.
Loutfi A, Coradeschi S, Saffiotti A:Maintaining coherent perceptual information using anchoring. Proc. of the 19th IJCAI Conf. 2005, Edinburgh: Morgan Kaufmann Publishers Inc,
Henson C, Sheth A, Thirunarayan K:Semantic perception: converting sensory observations to abstractions. IEEE Internet Comput. 2012, 16 (2): 26-34.
Thirunarayan K, Henson CA, Sheth AP:Situation awareness via abductive reasoning from semantic sensor data: a preliminary report. CTS. 2009, Washington, DC: IEEE Computer Society, 111-118.
Henson CA, Thirunarayan K, Sheth AP, Hitzler P:Representation of parsimonious covering theory in owl-dl. OWLED. CEUR Workshop Proceedings, Volume 796. Edited by: Dumontier M, Courtot M. 2011, France: CEUR-WS.org,
Samwald M, Freimuth RR, Luciano JS, Lin S, Powers RL, Marshall MS, Adlassnig K-P, Dumontier M, Boyce RD:An rdf/owl knowledge base for query answering and decision support in clinical pharmacogenetics. MedInfo. Studies in Health Technology and Informatics, Volume 192. 2013, IOS Press, 539-542.
Alirezaie M, Loutfi A:Automatic annotation of sensor data streams using abductive reasoning. Proceedings of the International Conference on Knowledge Engineering and Ontology Development (KEOD). 2013, SciTePress, [http://www.keod.ic3k.org/?y=2013],
Auer S, Lehmann J, Ngomo A-CN:Introduction to linked data and its lifecycle on the web. Proceedings of the 7th International Conference on Reasoning Web: Semantic Technologies for the Web of Data, RW’11. 2011, Berlin, Heidelberg: Springer-Verlag, 1-75.
Polleres A, Hogan A, Delbru R, Umbrich J:Rdfs and owl reasoning for linked data. Reasoning Web. 2013, Springer, 91-149.
Brewka G: Nonmonotonic Reasoning: Logical Foundations of Commonsense. 1991, Cambridge: Cambridge University Press, J. SIGART Bull
Cox PT, Pietrzykowski T:General diagnosis by abductive inference. SLP. 1987, Washington, DC: IEEE-CS, IEEE Computer Society, 183-189.
Bylander T, Allemang D, Tanner MC, Josephson JR:The computational complexity of abduction. Knowledge Representation, vol. 49. 1991, MIT Press, 25-60.
Graves GR, Rhodes PG:Tachycardia as a sign of early onset neonatal sepsis. Pediatr Infect Dis. 1984, 3 (5): 404-406. 10.1097/00006454-198409000-00002.
Clark AL, Sparrow JL, Coats AJ:Muscle fatigue and dyspnoea in chronic heart failure: two sides of the same coin?. Eur Heart J. 1995, 16 (1): 49-52. 10.1093/eurheartj/16.1.49.
Takrouri MSM:Intensive care unit. Int J Health. 2003,, ISPUB; 3.,
Compton M, Barnaghi P, Bermudez L, Garcia-Castro R, Corcho O, Cox S, Graybeal J, Hauswirth M, Henson C, Herzog A, Huang V, Janowicz K, Kelsey WD, Phuoc DL, Lefort L, Leggieri M, Neuhaus H, Nikolov A, Page K, Passant A, Sheth A, Taylor K:The ssn ontology of the w3c semantic sensor network incubator group. Web Semantics: Sci Serv Agents World Wide Web. 2012, 17: 25-32.
Kunder M: The size of the World Wide Web. [http://www.worldwidewebsize.com] last visit: 10 July 2014
de Marneffe M-C, MacCartney B, Manning CD:Generating typed dependency parses from phrase structure trees. LREC. 2006, Paris: ELRA/ELDA, 449-454.
Acknowledgements
This work has been supported by the Swedish National Research Council, Vetenskapsrådet, project nr 2010-4769, on cognitive olfaction.
The evaluation of the framework presented in this article has also been assisted by using the ICU data package provided for use in 1994 AI in Medicine symposium submissions [10].
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
MA: modelling and developing different parts of the framework. AL: supervising in modelling the framework and revising the manuscript. Both authors read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly credited. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Alirezaie, M., Loutfi, A. Automated reasoning using abduction for interpretation of medical signals. J Biomed Semant 5, 35 (2014). https://doi.org/10.1186/2041-1480-5-35
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/2041-1480-5-35

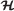 (set of all hypotheses) to a subset of
(set of all hypotheses) to a subset of  (set of manifestations) so that
(set of manifestations) so that  is a possible cause for . An accepted conclusion w.r.t the set-covering criterion is set such that .
is a possible cause for . An accepted conclusion w.r.t the set-covering criterion is set such that . as a subset of
as a subset of  is chosen as the solution set if for all other “covering” subsets of
is chosen as the solution set if for all other “covering” subsets of 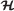 , namely
, namely 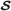 , .
, .







